Hi AFNI experts and users,
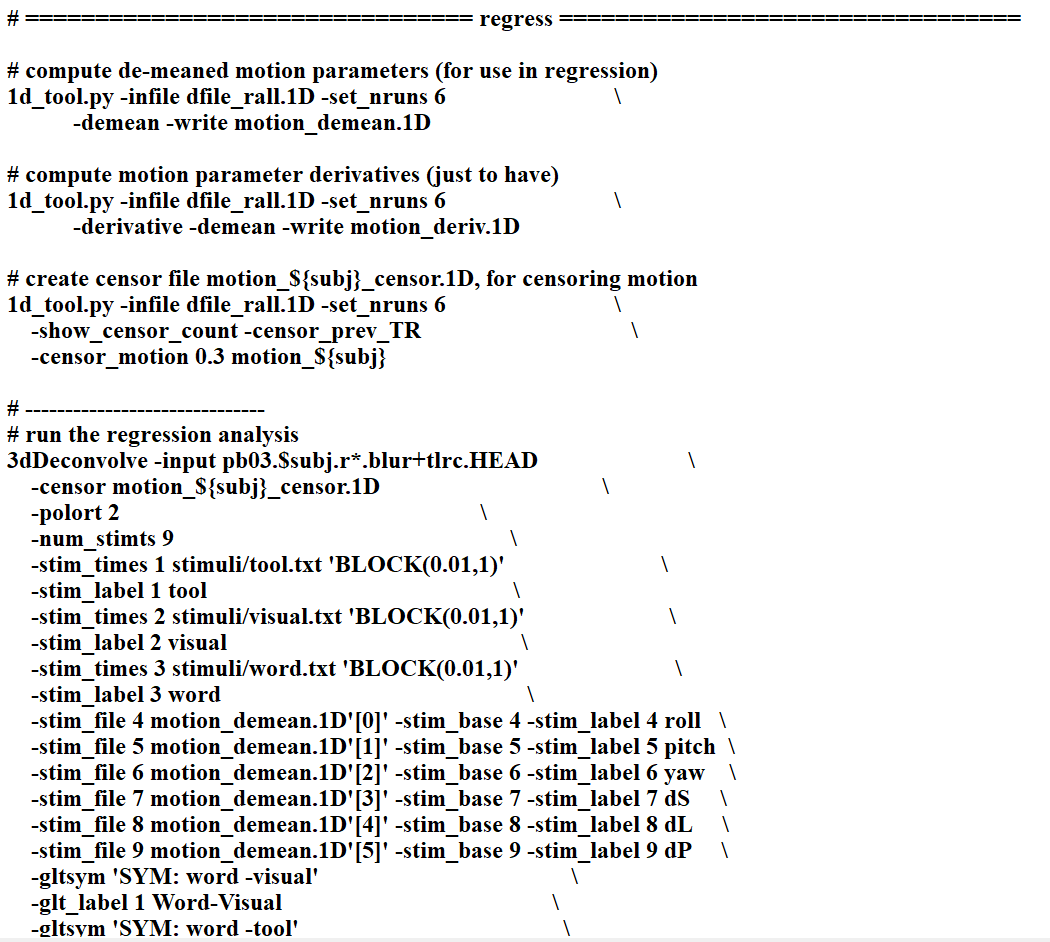
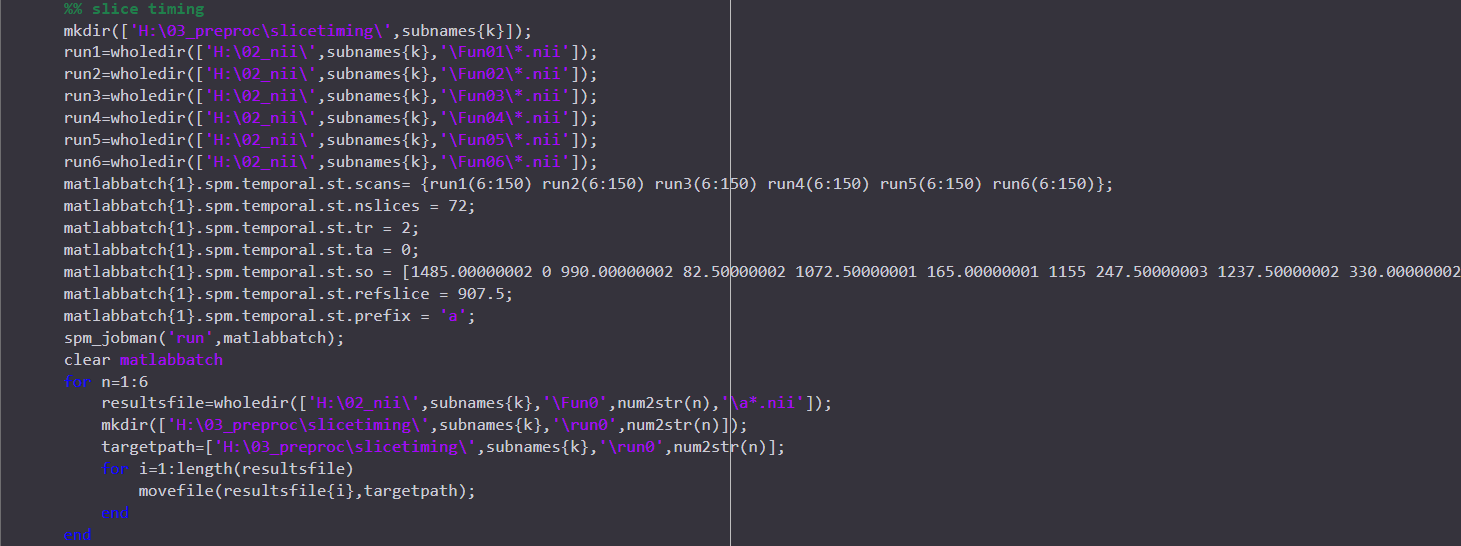
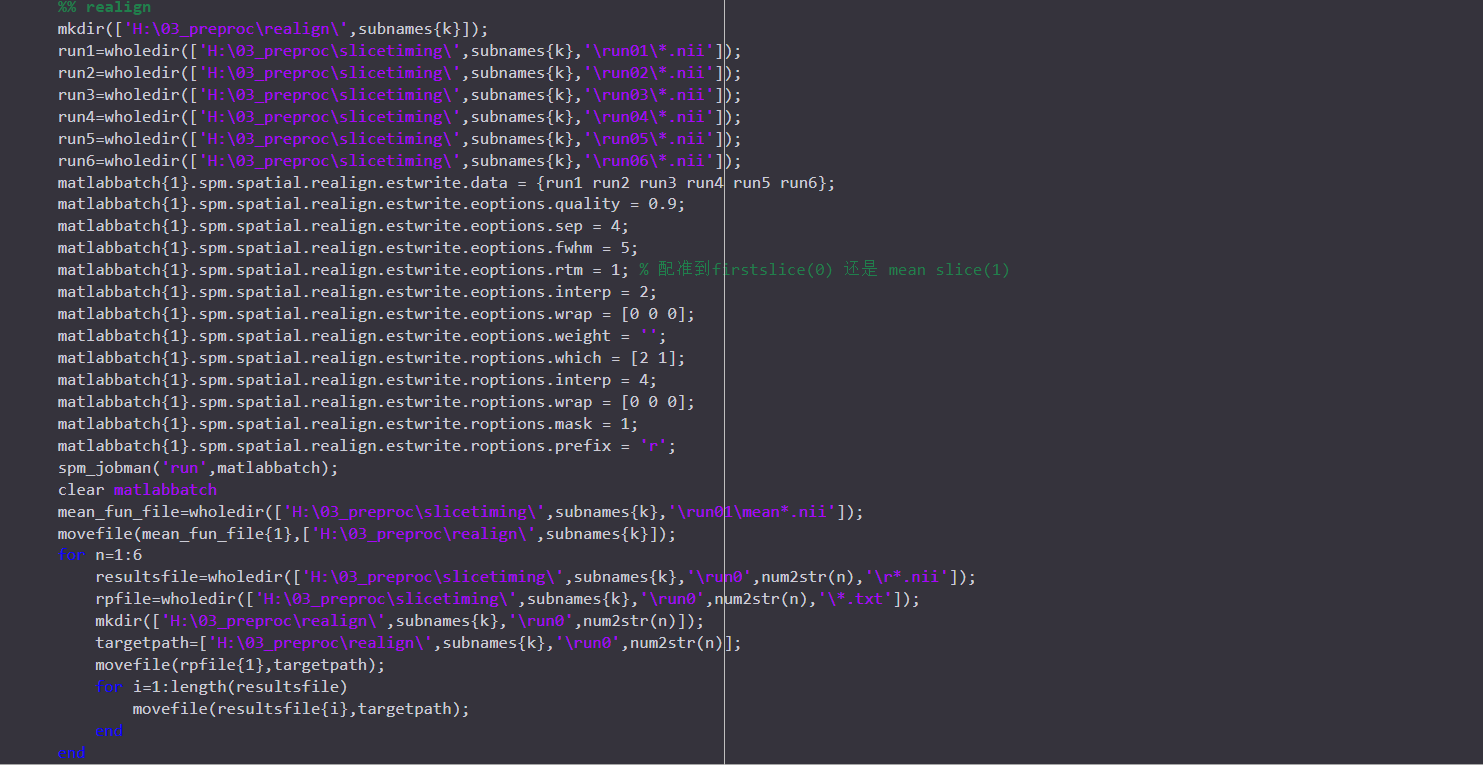
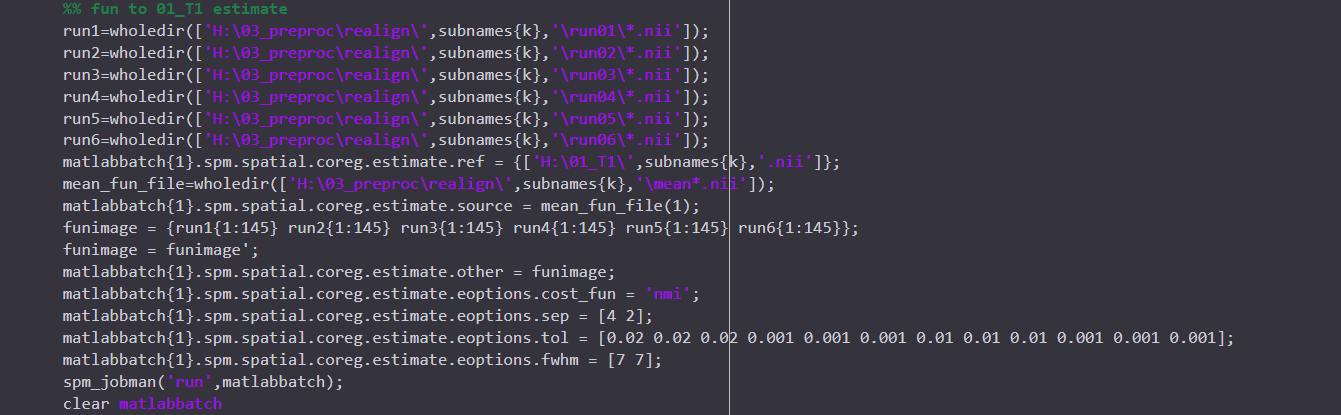
I just preprocessed my data using AFNI proc.py. And I also preprocessed the same data using SPM pepline before. But what made me confused is that the results from two toolbox are very different ..... This thing made me doubted the reproducibility and reliability of the analysis I have done, which are all based on the data after prrprocessing in SPM. I posted the preprocessed codes I used in two toolboxs here. Any information or help would be really appreciated! THANKSSS.
Hi-
To post code, it is better to copy+paste text and to put it within backticks, as described here. This will help us (and others) read it and/or copy+paste it ourselves, if need be.
To your question about processing, it would be better if you could please copy+paste your afni_proc.py command, so we can understand what processing pipeline you set up for FMRI with AFNI. That is the succinct form for understanding all your processing when using AFNI. I am not sure what the equivalent is with SPM, but if you could do something similar, that would be more helpful.
I haven't seen any of your results, so I don't know what your data is or how you are comparing things, which are key factors. I will just comment on a couple general points that might be useful.
Before doing any comparisons, there is the important step of doing quality control (QC), to make sure that the data are consistent and reasonable for your study goals, and that all processing steps completed successfully. This is described in more detail in the Open FMRI QC Project, for example, see here. There are several articles about QC'ing afni_proc.py results, such as Reynolds et al. 2023, Birn 2023, Lepping et al., 2023 and Teves et al. 2023. There was one group that used SPM, Di and Biswal, 2023, so you might be interested in that for evaluating your SPM data.
Comparing datasets or processing pipelines has a lot of considerations. The NARPS study was focused on this, where groups processed data using AFNI, FSL, SPM, fMRIprep and more. If comparisons come after thresholding, there can look like many differences in results. However, without thresholding inserted, results across the 60+ teams were actually very similar across the vast majority of teams, regardless of software, just with slightly different magnitude. This was shown in more detail in this paper, in which we looked at the NARPS teams results:
- Taylor PA, Reynolds RC, Calhoun V, Gonzalez-Castillo J, Handwerker DA, Bandettini PA, Mejia AF, Chen G (2023). Highlight Results, Don’t Hide Them: Enhance interpretation, reduce biases and improve reproducibility. Neuroimage 274:120138. doi: 10.1016/j.neuroimage.2023.120138
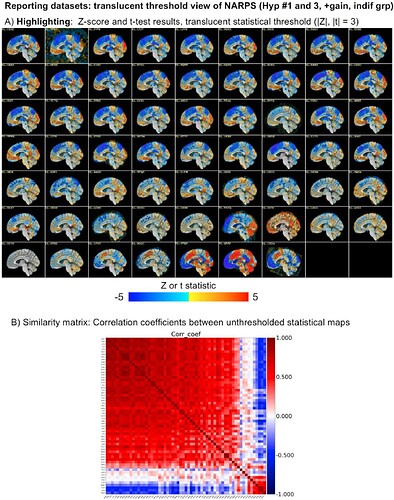
See for example Fig 8 from there, showing both the same slice across all subjects and the similarity matrix:
The vast majority of subjects have the similar positive/negative modeling pattern across the brain (big red square in the correlation matrix); the bottom red square is actually just teams that made a processing mistake or a different sign notation and have opposite sign results---but after sorting out that issue they are actually the same as the big red square group. So, in the end there are just a small fraction of very different results, which is not surprising.
You can check out other hypotheses and things from that paper---the results remain quite similar. The main less from this should be be careful how you do your meta analyses and how you look at your data. Inserting thresholding can/willl heavily bias you toward accentuating differences and reducing reproducibility. The NARPS study shows the challenges of meta analyses and how difficult/subtle they can be. The Highlight, Don't Hide paper/methodology shows simply showing more data in figures and reducing threshold-induced biases in comparisons leads to better understanding.
So, as you start looking at different pipelines, it is good to keep in mind both the need for quality control to check data+processing. Then also please keep in mind the question of how to do fair comparisons that are not biased from the start.
--pt
If you wanted to add some useful images - when you say they are different what do you mean? Maybe an image of the same area at the same threshold in both programs would help.
Additionally, sharing the design matricies from each software might help - this is X.jpg in AFNI (in results folder), and I think you can capture it from the Results view in SPM.
I am familiar with SPM, but it has been a while. For you events ("word", "tool", "visual", it looks like they are at 0, 2, 4 seconds, and then 14, 20, 32, and so on and so forth, but they all last 0 seconds? is that right? Duration is zero in SPM, and 0.01 in AFNI, which feel like strange values for a stimulus event. I am having a hard time understanding what this experiment was like, so a brief description might also help.
Hi!ptaylor,
Thanks so much that your kindful relpy.
I just attend to do the quality control to my data. But I found that there is no QC files in the results in my resultsfiles. And I found that may because there is no apqc.py in my afni programs. Could you please tell me how can i generate the QC files in my preprogress ?
Thanks again!
hi, dowdlelt
thanks for your kindful reply!
the 0.1 in BLOCK(0.1,1) is becuse that the design of my exp is Event_Related(ER), and the afni document said that if ER design, i should change the duration paramater to a very small value.
And for the same reason, but in spm, this value should set as 0.
Hi-
Can you please copy+paste the text output by this command, which will provide a lot of useful information about your current AFNI and system setup:
afni_system_check.py -check_all
? It is possible you have an old version of AFNI (though it would have to be very old not to make the QC directory), or missing dependencies.
Also, the program to look for is open_apqc.py to open the QC files, and apqc_make*.py to create the QC.
--pt
sure
[hanlab@cu72 ~]$ afni_system_check.py -check_all
-------------------------------- general ---------------------------------
architecture: 64bit ELF
system: Linux
release: 2.6.32-431.el6.x86_64
version: #1 SMP Fri Nov 22 03:15:09 UTC 2013
distribution: CentOS 6.5 Final
number of CPUs: 64
apparent login shell: bash
--------------------- AFNI and related program tests ---------------------
which afni : /opt/software/linux_xorg7_64/afni
afni version : Precompiled binary linux_xorg7_64: Aug 12 2015
which python : /usr/bin/python
python version : 2.6.6
which R : /usr/bin/R
R version : R version 3.4.2 (2017-09-28) -- "Short Summer"
which tcsh : /bin/tcsh
instances of various programs found in PATH:
afni : 1 (/opt/software/linux_xorg7_64/afni)
R : 1 (/usr/bin/R)
python : 2
/usr/bin/python
/opt/software/anaconda3/bin/python3.6
testing ability to start various programs...
afni : success
suma : success
3dSkullStrip : success
uber_subject.py : success
3dAllineate : success
3dRSFC : success
SurfMesh : success
checking for $HOME files...
.afnirc : missing
.sumarc : missing
.afni/help/all_progs.COMP : missing
------------------------------ python libs -------------------------------
++ module 'PyQt4' found at /usr/lib64/python2.6/site-packages/PyQt4
++ module loaded: PyQt4
------------------------------- path vars --------------------------------
PATH = /opt/software/MATLAB/R2015b/bin/:/opt/software/fsl/bin:/opt/intel//impi/5.0.2.044/intel64/bin:/opt/intel/composer_xe_2015.1.133/bin/intel64:/opt/intel/composer_xe_2015.1.133/debugger/gdb/intel64_mic/bin:/opt/software/tracula.update.centos6_x86_64.5.3.2014_05_26:/opt/software/mrtrix3/bin/:/opt/software/freesurfer-Linux-centos6_x86_64-stable-pub-v5.3.0/bin:/opt/software/freesurfer-Linux-centos6_x86_64-stable-pub-v5.3.0/bin:/opt/software/freesurfer-Linux-centos6_x86_64-stable-pub-v5.3.0/fsfast/bin:/opt/software/freesurfer-Linux-centos6_x86_64-stable-pub-v5.3.0/tktools:/opt/software/freesurfer-Linux-centos6_x86_64-stable-pub-v5.3.0/mni/bin:/opt/software/mricron/:/opt/software/linux_xorg7_64/:/opt/software/DTK_trackvis/:/opt/software/MATLAB/R2017a/bin/:/usr/local/maui/sbin:/usr/local/maui/bin:/opt/tsce/share/bin:/opt/tsce/share/sbin:/opt/tsce/share/jdk1.6.0_10/bin:/opt/tsce/share/jdk1.6.0_10/jre/bin:/usr/lib64/qt-3.3/bin:/usr/kerberos/sbin:/usr/kerberos/bin:/usr/local/bin:/bin:/usr/bin:/usr/local/sbin:/usr/sbin:/sbin:/opt/ibutils/bin:.:/opt/software/anaconda3/bin:/brain/hanlab/bin
PYTHONPATH =
LD_LIBRARY_PATH = /opt/intel//impi/5.0.2.044/intel64/lib:/opt/intel/composer_xe_2015.1.133/compiler/lib/intel64:/opt/intel/composer_xe_2015.1.133/mpirt/lib/intel64:/opt/intel/composer_xe_2015.1.133/ipp/../compiler/lib/intel64:/opt/intel/composer_xe_2015.1.133/ipp/lib/intel64:/opt/intel/composer_xe_2015.1.133/compiler/lib/intel64:/opt/intel/composer_xe_2015.1.133/mkl/lib/intel64:/opt/intel/composer_xe_2015.1.133/tbb/lib/intel64/gcc4.4
DYLD_LIBRARY_PATH =
DYLD_FALLBACK_LIBRARY_PATH =
------------------------------ data checks -------------------------------
data dir : missing AFNI_data6
data dir : missing suma_demo
data dir : missing FATCAT_DEMO
atlas : found TT_N27+tlrc under /opt/software/linux_xorg7_64/
------------------------------ OS specific -------------------------------
---------------------------- general comments ----------------------------
* consider copying AFNI.afnirc to ~/.afnirc
* consider running "suma -update_env" for .sumarc
* consider running "apsearch -update_all_afni_help"
yes, the version of afni in our server is related in 2015, I think it is old
I asked our server administrator, the later version of afni required python 3, but in our server, we used python 2.7. Is it possible to use later version of afni in this environment?
Howdy-
It appears that the AFNI version you have is from 2015:
Thaaaaaaaaaat is quite, quite old. There have been so many updates to programs and new functionalities added since then. I can only think you would have to update it.
Most functionality in AFNI runs on both Python 2.* and 3.*, actually, despite the challenges that presents. However, you should be aware that Python 2.* has officially been deprecated, meaning that it is no longer actively maintained. Many pieces of software will only really work on Python 3.*, and so you might want to consider asking your local server administrator to update that, as well.
--pt
ok, got it. thank you so much!
Lot of Information Found,'Thanks