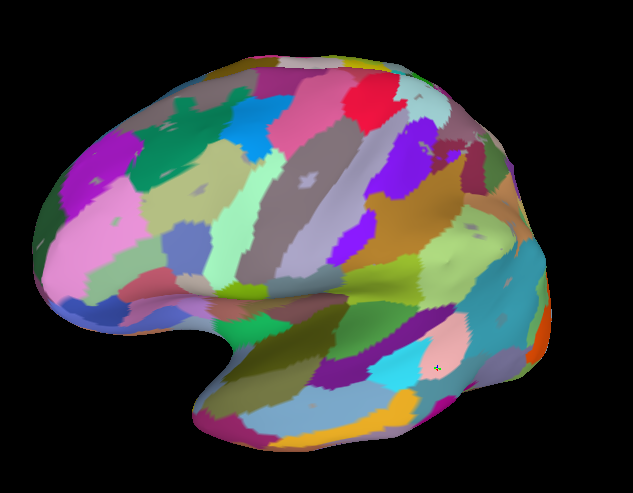
How are you mapping the data to the surface? If you are using the default afni-suma communications, then you will only get the intersections of the surface with the voxels of the volume. That will often leave holes if the mask bends even just a little away from the volume. If you are doing this interactively, you can use the Vol2surf plugin to adjust how that’s done and allow for the mode/median/maximum along the segments from the pial to smoothwm surfaces. The mode usually makes the most sense, I think.
The Brainnetome atlas was based on the HCP-40 dataset, a templatized subset of the Human Connectome Project anatomical data. It’s aligned to 2009c dataset, but it is slightly different, so it doesn’t fill the edges perfectly either.
To fix both problems, you can use the interactive Vol2Surf plugin or use this script to project the atlas to the surface or download the niml.dset mapping here. Here I project with 3dVol2Surf using the non-zero mode, and then smooth the atlas on the surface with modal smoothing in a 5mm neighborhood. Find the links below for the atlas projection data and the MNI surfaces. The projected data is not perfectly filled and smooth, but it is usable.
https://afni.nimh.nih.gov/pub/dist/atlases/brainnetome/bn_surfs.tgz
Viewing the surface atlas, now in a surface mapping file that ends with .niml.dset, involves loading that dataset in suma.
- Start suma with directory of spec file for the MNI surfaces, here with both hemispheres
suma -spec ~/suma_MNI152_2009/std.60.MNI152_2009_both.spec
-sv ~/abin/MNI152_2009_template.nii.gz
- Right click on a surface and Ctrl-s to open the surface object controller.
- Load dset in the lower left part of that menu.
- Select the bnsurfatlas_beta0_mode5mm.${hemi}.niml.dset depending on which hemisphere surface is selected. The contralateral hemisphere will automatically be loaded.
- Change the colormap “Cmp” to be ROI_glasbey_256 or a similar colormap made for ROI indices.
- Adjust the shading by turning off background convexity coloring with the ‘b’ key over the suma image viewer/
#!/bin/tcsh
project Brainnetome atlas to the MNI 152 2009c surface
this assumes the HCP40 is close enough to the 2009c asymmetric volume
where MNI152 2009c surfaces are
set surfpath = ~/suma_MNI152_2009
where the BN atlas is
set atlaspath = ~/abin
for both left and right hemispheres
foreach hemi ( lh rh )
project from the volume to the surface
use the non-zero mode to fill surface with something if any voxel along the path
has an atlas voxel
using the ld60 low resolution surface, but you can use ld141 with std.141…
3dVol2Surf -spec surfpath/std.60.MNI152_2009_both.spec \
-surf_A std.60.{hemi}.smoothwm.gii
-surf_B std.60.${hemi}.pial.gii -sv $atlaspath/MNI152_2009_template.nii.gz
-grid_parent $atlaspath/BN_Atlas_246_1mm.nii.gz -gp_index 0
-map_func nzmode -f_steps 10 -f_index nodes
-cmask "-a atlaspath/BN_Atlas_246_1mm.nii.gz -expr astep(a,0.000000)" \
-out_niml bnsurfatlas_beta0.{hemi}.niml.dset
this is a bonus set… the projection is quite noisy, so we can use the mode
again, but this time on the surface with a 5mm neighborhood
SurfLocalstat -hood 5 -stat mode -input bnsurfatlas_beta0.${hemi}.niml.dset
-i_gii surfpath/std.60.{hemi}.smoothwm.gii
-prefix bnsurfatlas_beta0_mode5mm.${hemi}.niml.dset
end
The MNI 2009c asymmetric surfaces are here:
https://afni.nimh.nih.gov/pub/dist/tgz/suma_MNI152_2009.tgz
While not exactly what you asked for, Brainnetome has a surface workflow that transforms atlas to the subject’s surface (BN_Atlas_freesurfer.zip)
http://atlas.brainnetome.org/download.html