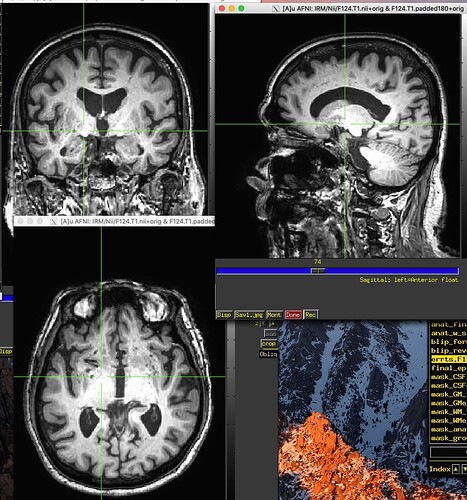
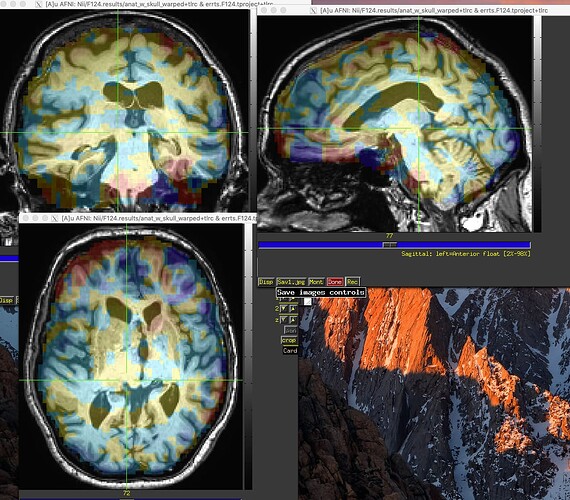
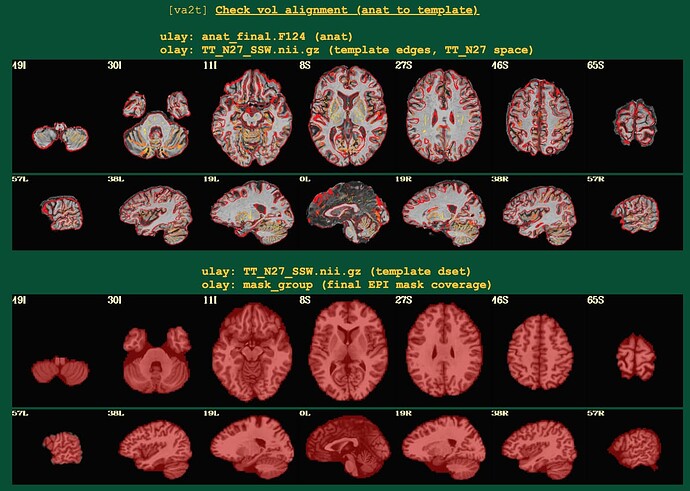
Here's the output. The 3dMVM failure is becasue of the version of Mac. I asked here before.
-------------------------------- general ---------------------------------
architecture: 64bit
cpu type: i386
system: Darwin
release: 18.7.0
version: Darwin Kernel Version 18.7.0: Tue Jun 22 19:37:08 PDT 2021; root:xnu-4903.278.70~1/RELEASE_X86_64
distribution: 10.14.6
number of CPUs: 8
apparent login shell: bash
shell RC file: .bashrc (exists)
--------------------- AFNI and related program tests ---------------------
which afni : /Users/xiyuezh/abin/afni
afni version : Precompiled binary macos_10.12_local: May 30 2023
: AFNI_23.1.07 'Publius Helvius Pertinax'
AFNI_version.txt : AFNI_23.1.07, macos_10.12_local, May 30 2023
which python : /usr/bin/python
python version : 2.7.16
which R : /usr/local/bin/R
R version : R version 4.2.2 (2022-10-31) -- "Innocent and Trusting"
which tcsh : /bin/tcsh
instances of various programs found in PATH:
afni : 1 (/Users/xiyuezh/abin/afni)
R : 1 (/Library/Frameworks/R.framework/Versions/4.2/Resources/bin/R)
python : 1 (/usr/bin/python)
python2 : 0
python3 : 1 (/Library/Frameworks/Python.framework/Versions/3.9/bin/python3.9)
Xvfb : 1 (/opt/X11/bin/Xvfb)
** have python3 but not python2
testing ability to start various programs...
afni : success
suma : success
3dSkullStrip : success
uber_subject.py : success
3dAllineate : success
3dRSFC : success
SurfMesh : success
3dClustSim : success
3dMVM : FAILURE
Error in dyn.load(ll) :
unable to load shared object '/Users/xiyuezh/abin/R_io.so':
dlopen(/Users/xiyuezh/abin/R_io.so, 6): Library not loaded: /sw/Library/Frameworks/R.framework/Versions/3.6/Resources/lib/libR.dylib
Referenced from: /Users/xiyuezh/abin/R_io.so
Reason: image not found
Calls: source ... withVisible -> eval -> eval -> set_R_io -> dyn.load
Execution halted
checking for R packages...
rPkgsInstall -pkgs ALL -check : success
R RHOME : /Library/Frameworks/R.framework/Resources
checking for $HOME files...
.afnirc : found
.sumarc : found
.afni/help/all_progs.COMP : found
------------------------------ python libs -------------------------------
** failed to load module PyQt4
-- PyQt4 is no longer needed for an AFNI bootcamp
++ module loaded: matplotlib.pyplot
module file : /System/Library/Frameworks/Python.framework/Versions/2.7/Extras/lib/python/matplotlib/pyplot.pyc
matplotlib version : 1.3.1
** need maptplotlib version 2.2+ for APQC
-- python binaries under /sw/bin:
/sw/bin/python2.7
-- python binaries under /usr/local/bin:
/usr/local/bin/python3 (sym link to /Library/Frameworks/Python.framework/Versions/3.9/bin/python3.9)
/usr/local/bin/python3.9 (sym link to /Library/Frameworks/Python.framework/Versions/3.9/bin/python3.9)
-------------------------------- env vars --------------------------------
PATH = /usr/local/fsl/bin:/opt/anaconda3/bin:/Library/Frameworks/Python.framework/Versions/3.9/bin:/usr/local/bin:/usr/bin:/bin:/usr/sbin:/sbin:/opt/X11/bin:/Users/xiyuezh/abin
PYTHONPATH =
R_LIBS =
LD_LIBRARY_PATH =
DYLD_LIBRARY_PATH (sub-shell) = :/opt/X11/lib/flat_namespace:/opt/X11/lib/flat_namespace
DYLD_FALLBACK_LIBRARY_PATH (sub-shell) =
----------------------------- eval dot files -----------------------------
** warning: .tcshrc does NOT seem to contain 'source .cshrc'
-- (csh and tcsh will use different files)
-- considered operations: path, flatdir, apsearch
-- note: followers should not need edits, so edit flags should be 0
(have 0 follower(s), which can be ignored)
dot file test : want 3 modifications across 3 files:
file path flatdir apsearch follower
--------------- ---- ------- -------- --------
.tcshrc 1 1 1 0
.cshrc 0 0 0 0
.bashrc 0 0 0 0
------------------------------ data checks -------------------------------
data dir : found AFNI_data6 under $HOME (2976900M Avail)
top history: 20 Feb 2020 [rickr]: updated FT_analysis examples
data dir : found AFNI_demos under $HOME
top history: ...ct 2020 [taylorp]: updated scripts under FATCAT_DEMO
data dir : found suma_demo under $HOME
top history: ...s_New/data/Build_tmp on Mon Mar 4 11:56:45 EST 2013
data dir : found afni_handouts under $HOME
atlas : found TT_N27+tlrc under /Users/xiyuezh/abin
------------------------------ OS specific -------------------------------
XQuartz version : 2.8.1
which brew : /usr/local/bin/brew
brew version : Homebrew 4.0.22
++ found PyQt4 under /sw/lib/qt4-mac/lib/python2.7/site-packages
** seem to be using fink python2.7 but need python
consider: sudo ln -s /sw/bin/python2.7 /sw/bin/python
** warning: have fink PyQt4, but non-fink python /usr/bin/python
++ found 1 dylib files under '/opt/X11/lib/flat_namespace'
-- found 'libXt' dylib files:
/opt/X11/lib/flat_namespace/libXt.6.dylib
-- recent OS X, cheating to check DYLD_LIBRARY_PATH in cur shell 'bash'...
++ found evar DYLD_LIBRARY_PATH = :/opt/X11/lib/flat_namespace:/opt/X11/lib/flat_namespace
-- recent OS X, cheating to check DYLD_LIBRARY_PATH in shell 'tcsh'...
** env var DYLD_LIBRARY_PATH not set to contain /opt/X11/lib/flat_namespace
========================= summary, please fix: =========================
- just be aware: login shell 'bash', but our code examples use 'tcsh'
- AFNI programs show FAILURE
- check for partial install of PyQt4
- dot file test : want 3 modifications across 3 files:
- consider installing gcc under homebrew
- consider installing glib under homebrew
- please set DYLD_LIBRARY_PATH to /opt/X11/lib/flat_namespace in tcsh