Hi AFNI gurus,
I'm working with a sample of about 200 and am using SSwarper for both our task-based and structural analyses (I have found that SSwarper works better than FreeSurfer on my data for the initial skullstripping).
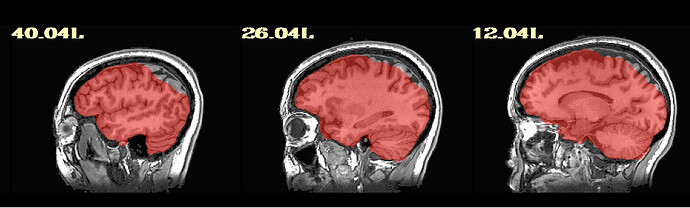
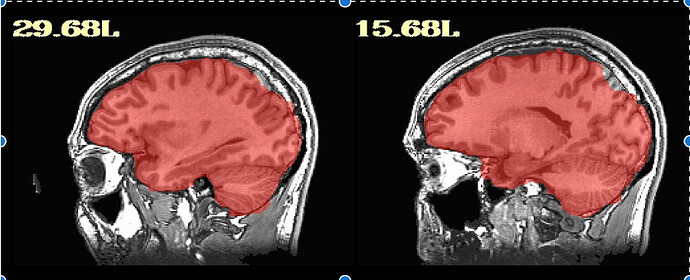
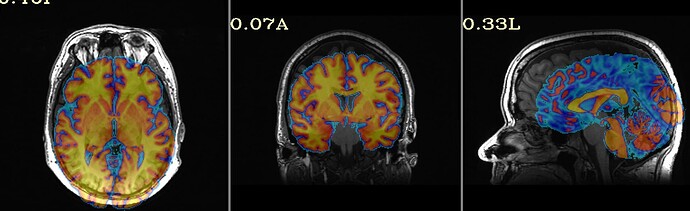
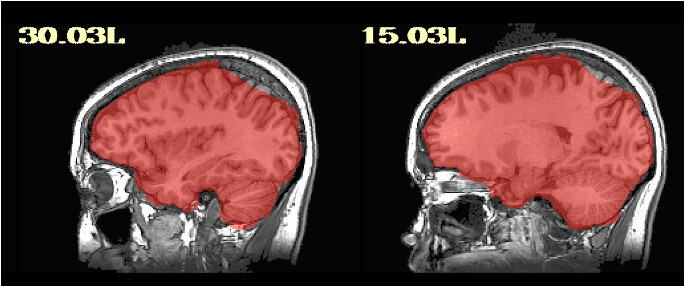
For the majority of participants, the output is really good; however, for about 20, there are some issues. They are usually in the superior parietal region, which makes me think something could be adjusted for those participants. I am running Align_Centers prior to sswarper and have played around with a couple of the parameters (lpa, giant_move) without much luck. I'm wondering if anyone has further suggestions! Of note, it is a sample of heavy drinkers, so some of the brains display atrophy/ enlarged sulci.
AFNI version info (afni -ver): AFNI_22.3.07
code text # or delete if not needed
Howdy-
We do have an updated version of @SSwarper, called sswarper2. It might help with those subjects. It has very similar syntax and outputs to @SSwarper2, so it should be directly swappable in your pipeline. It also has more QC outputs and history saved (to help troubleshooting).
You would need a more recent AFNI version to have it, since we added this program in early 2024.
Maybe a side note: giant_move would probably be most useful in cases of catastrophic skullstripping failure, to help overcome initial woe with relative coordinates. If things are "pretty good, but locally not great", then I wouldn't expect that to have much difference. Probably it is good that Align_Centers was run to start, esp. if coordinates for the input anatomical didn't have a well-centered coordinate origin.
--pt
1 Like
Hi there,
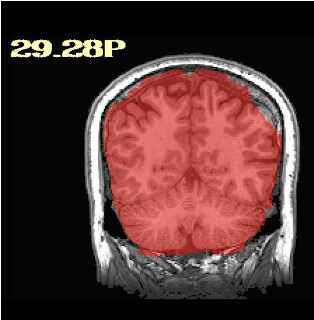
Thanks so much for getting back to me. I did as you said an have rerun these participants with sswarper2. Unfortunately, it did not seem to resolve any of the issues. As you mentioned, I do now have the folder with additional files, although I'm not sure how best to use them. The initial alignment is slightly off in most of the participants (see below) - is this normal? Do you have any further recommendations regarding the additions to the sswarper2 command that may be worth trying?
Thanks so much for your time,
Carly
Sorry, I let this thread slip from my attention.
What are some example images of the skullstripping output, like the QC_anatSS*.jpg images from sswarper2?
--pt