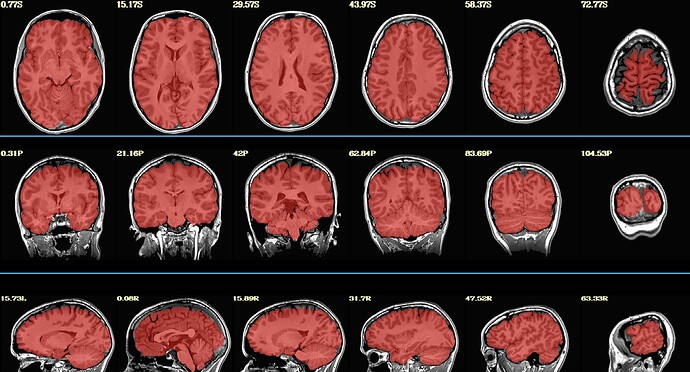
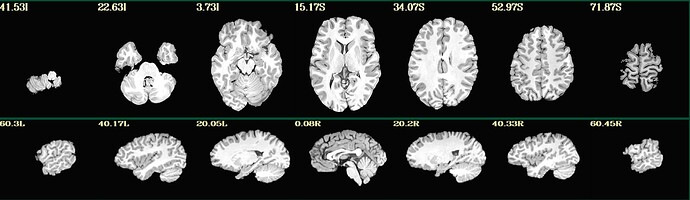
Hello, we have a few participants where parts of their brain got stripped along with their skull (see example below). They were originally processed on sswarper, and sswarper2 / push to edge both made things worse.
We ended up re-running sswarper2 locally using AFNI version 26.0.05. With this, the original and warped anatomical look ok in afni, but the jpgs in sswaper2 and QC html files still look aggressively stripped. What is the recommended procedure for attempting to fix this? Thank you!
Howdy-
Sorry for the delay in replying, Bootcamping took my mental energy this week.
The JPGs and the data views should be identical, assuming there is no obliquity in the inputs.
For tweaking the results, since overall the results are good and there are just some edge regions to adjust, I would consider changing the final cost function used for the nonlinear parts. The default cost function for the final nonlinear stage is "pcl", but you could adjust this to "lpa" with:
-cost_nl_final lpa
I would try either "lpa" or "nmi" for the final cost there.
--pt