AFNI version info (afni -ver): 25.1.04
Halo everyone,
thanks a lot for a wonderful toolbox and for being such a supportive community! ![]()
We have a problem with our 7T single-echo partial volume dataset, and were hoping you might help us solve it.
In brief, we collected three runs of single-echo multi-band partial volume data (focusing on the amygdala), as well as SBRefs and AP/PA scans for each run, and one mp2rage for the whole session. We want to analyze each individual separately without warping the images to standardized space.
Before running afni_proc (see below), we 'cleaned' the combined mp2rage scan using the INV2 scan together with the MPRAGEise python script. All looks well.
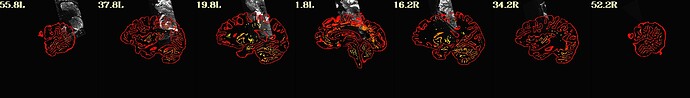
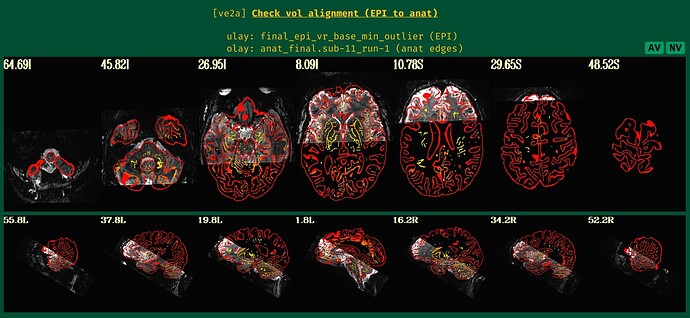
However, for some subjects we get very bad realignments after afni_proc:
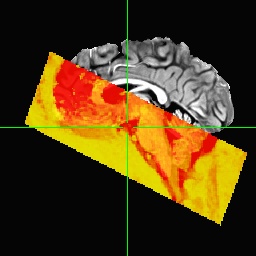
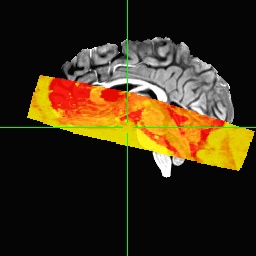
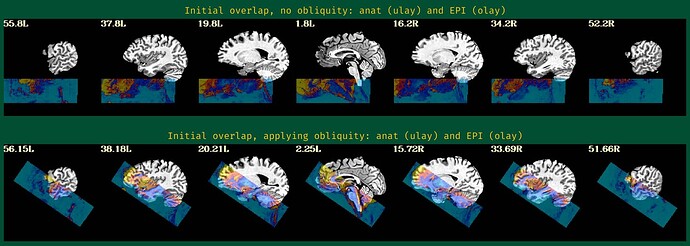
We tried to realign the anatomical to the epis using align_epi_anat prior to running afni_proc, which seems to work well, i.e. the anatomical looks nicely aligned to the epis. However, when using this realigned anatomical as input to afni_proc, we get very bad results again (sorry cannot paste more embedded media here because new user):
We feel like we are missing something basic (first time AFNI users), and would be very grateful for any advice!
Thanks a lot and have a nice day!
Kristoffer
afni_proc.py \
-subj_id ${subj}_${run} \
-out_dir ${parrun_outdir} \
-script ${scripts_dir}/${script_file} \
-dsets ${epis} \
-copy_anat ${anat} \
-anat_has_skull no \
-volreg_base_dset ${SBref} \
-align_epi_ext_dset ${SBref} \
-blocks despike tshift align volreg mask scale regress \
-anat_follower anat_w_skull anat ${anat_with_skull} \
-anat_follower_ROI allROIs_a anat ${suma_dir}/aparc.a2009s+aseg_REN_all.nii.gz \
-anat_follower_ROI allROIs_epi epi ${suma_dir}/aparc.a2009s+aseg_REN_all.nii.gz \
-anat_follower_ROI L_LatNuc_a anat ${L_LatNuc} \
-anat_follower_ROI L_BasNuc_a anat ${L_BasNuc} \
-anat_follower_ROI L_CenNuc_a anat ${L_CenNuc} \
-anat_follower_ROI L_MedNuc_a anat ${L_MedNuc} \
-anat_follower_ROI L_CorNuc_a anat ${L_CorNuc} \
-anat_follower_ROI L_AccBasNuc_a anat ${L_AccBasNuc} \
-anat_follower_ROI L_CATA_a anat ${L_CATA} \
-anat_follower_ROI L_AAA_a anat ${L_AAA} \
-anat_follower_ROI L_ParNuc_a anat ${L_ParNuc} \
-anat_follower_ROI R_LatNuc_a anat ${R_LatNuc} \
-anat_follower_ROI R_BasNuc_a anat ${R_BasNuc} \
-anat_follower_ROI R_CenNuc_a anat ${R_CenNuc} \
-anat_follower_ROI R_MedNuc_a anat ${R_MedNuc} \
-anat_follower_ROI R_CorNuc_a anat ${R_CorNuc} \
-anat_follower_ROI R_AccBasNuc_a anat ${R_AccBasNuc} \
-anat_follower_ROI R_CATA_a anat ${R_CATA} \
-anat_follower_ROI R_AAA_a anat ${R_AAA} \
-anat_follower_ROI R_ParNuc_a anat ${R_ParNuc} \
-anat_follower_ROI L_LatNuc_epi epi ${L_LatNuc} \
-anat_follower_ROI L_BasNuc_epi epi ${L_BasNuc} \
-anat_follower_ROI L_CenNuc_epi epi ${L_CenNuc} \
-anat_follower_ROI L_MedNuc_epi epi ${L_MedNuc} \
-anat_follower_ROI L_CorNuc_epi epi ${L_CorNuc} \
-anat_follower_ROI L_AccBasNuc_epi epi ${L_AccBasNuc} \
-anat_follower_ROI L_CATA_epi epi ${L_CATA} \
-anat_follower_ROI L_AAA_epi epi ${L_AAA} \
-anat_follower_ROI L_ParNuc_epi epi ${L_ParNuc} \
-anat_follower_ROI R_LatNuc_epi epi ${R_LatNuc} \
-anat_follower_ROI R_BasNuc_epi epi ${R_BasNuc} \
-anat_follower_ROI R_CenNuc_epi epi ${R_CenNuc} \
-anat_follower_ROI R_MedNuc_epi epi ${R_MedNuc} \
-anat_follower_ROI R_CorNuc_epi epi ${R_CorNuc} \
-anat_follower_ROI R_AccBasNuc_epi epi ${R_AccBasNuc} \
-anat_follower_ROI R_CATA_epi epi ${R_CATA} \
-anat_follower_ROI R_AAA_epi epi ${R_AAA} \
-anat_follower_ROI R_ParNuc_epi epi ${R_ParNuc} \
-tshift_interp -wsinc9 \
-align_unifize_epi local \
-align_opts_aea -big_move -check_flip -cost lpc+ZZ \
-mask_epi_anat yes \
-volreg_compute_tsnr yes \
-volreg_align_e2a \
-regress_compute_tsnr_stats L_LatNuc_epi 1\
-regress_compute_tsnr_stats R_LatNuc_epi 1\
-regress_compute_tsnr_stats L_BasNuc_epi 1\
-regress_compute_tsnr_stats R_BasNuc_epi 1\
-regress_compute_tsnr_stats L_CenNuc_epi 1\
-regress_compute_tsnr_stats R_CenNuc_epi 1\
-regress_compute_tsnr_stats L_MedNuc_epi 1\
-regress_compute_tsnr_stats R_MedNuc_epi 1\
-regress_compute_tsnr_stats L_CorNuc_epi 1\
-regress_compute_tsnr_stats R_CorNuc_epi 1\
-regress_compute_tsnr_stats allROIs_epi 17 33 \
-regress_apply_mot_types demean deriv \
-regress_censor_motion 0.3 \
-regress_censor_outliers 0.05 \
-regress_motion_per_run \
-regress_compute_fitts \
-regress_make_ideal_sum sum_ideal.1D \
-regress_reml_exec \
-regress_run_clustsim no \
-html_review_style pythonic \
-execute