Hi,
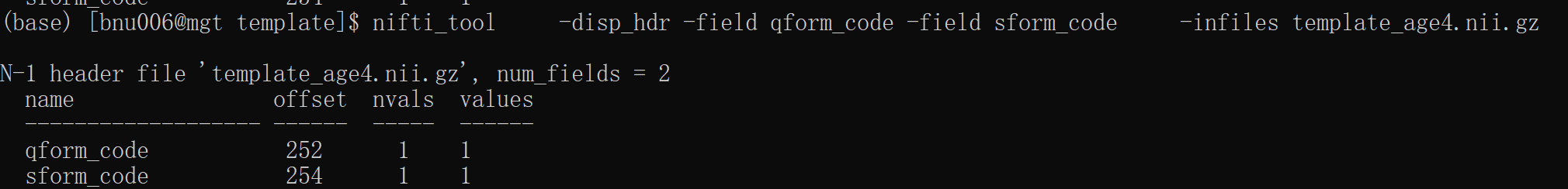
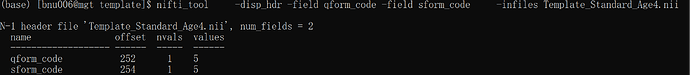
I have created my specific age template using ants and then registered T1 to this template with @animal_warper command. However, I failed running afni_proc.py with the following errors:
The code I used with afni_proc.py was:
set dsets_NL_warp = (/data/home/bnu006/UW-Madison_Rhesus_MRI/preprocess/data_13_aw/sub-2210/sub-2210_anat_warp2std_nsu.nii.gz \
/data/home/bnu006/UW-Madison_Rhesus_MRI/preprocess/data_13_aw/sub-2210/sub-2210_anat_composite_linear_to_template.1D \
/data/home/bnu006/UW-Madison_Rhesus_MRI/preprocess/data_13_aw/sub-2210/sub-2210_anat_shft_WARP.nii.gz )
afni_proc.py \
-subj_id sub-2210 \
-blocks tshift align tlrc volreg blur mask scale regress \
-dsets /data/home/bnu006/UW-Madison_Rhesus_MRI/preprocess/try_one/sub-2210/func/sub-2210_task-rest_bold.nii.gz \
-copy_anat /data/home/bnu006/UW-Madison_Rhesus_MRI/preprocess/data_13_aw/sub-2210/sub-2210_anat_nsu.nii.gz \
-anat_has_skull no \
-anat_uniform_method none \
-radial_correlate_blocks tcat volreg regress \
-radial_correlate_opts -sphere_rad 14 \
-tcat_remove_first_trs 10 \
-volreg_align_to MIN_OUTLIER \
-volreg_align_e2a \
-volreg_warp_master /data/home/bnu006/UW-Madison_Rhesus_MRI/preprocess/data_age4_basic/template/template_age4.nii.gz \
-volreg_tlrc_warp \
-volreg_compute_tsnr yes \
-align_opts_aea -cost nmi -epi_strip 3dSkullStrip -skullstrip_opts -monkey \
-check_flip -feature_size 0.5 \
-align_unifize_epi local \
-tlrc_base /data/home/bnu006/UW-Madison_Rhesus_MRI/preprocess/data_age4_basic/template/template_age4.nii.gz \
-tlrc_NL_warp \
-tlrc_NL_warped_dsets ${dsets_NL_warp} \
-blur_size 2.5 \
-regress_motion_per_run \
-regress_apply_mot_types demean deriv \
-regress_polort 2 \
-regress_bandpass 0.01 0.1 \
-regress_est_blur_errts \
-regress_est_blur_epits \
-regress_run_clustsim no \
-html_review_style pythonic
AFNI version info (afni -ver): Precompiled binary linux_centos_7_64: Feb 28 2024 (Version AFNI_24.0.09 'Caracalla')
Thanks,
Ruilin