Hi-
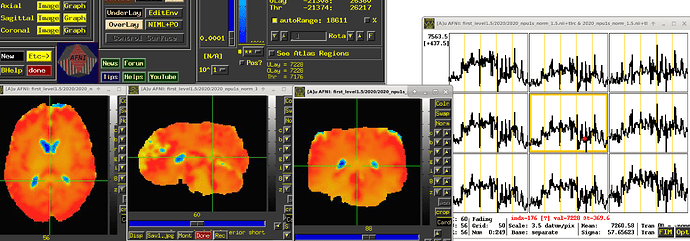
The EPI data acquired on MRI scanners for FMRI studies does not have meaningful, interpretable units. It is merely a blood oxygenation level dependent (BOLD) signal whose changes we can see over time, and does not have inherent physical interpretation. Even on the same scanner, different subjects' BOLD time series might have very different numerical baseline values, and we cannot directly compare values across them---a value of "500" has no inherent meaning, because the baseline for one subject might be 1000 and for another 478, and that 500 represents something pretty different in each case.
Consider a regression model:
y= \beta_0 + \beta_1 x_1 +\beta_2 x_2+ ... \epsilon
where y is the input data, each x_i is a regressor we put in and each \beta_i is the associated coefficient that we will solve for in the model (and the \epsilon is the residuals or leftover stuff from the input that wasn't fit by the combo of regressors); when we solve the GLM we get a set of estimated coefficient values, \hat{\beta}_i and for each one an associated statistic, t_i (which is essentially a number that says how many standard errors \sigma_i = \hat{\beta}_i/t_i the signal is away from zero).
From a "units" point of view, each \beta_i (or equivalently \hat{\beta}_i) will have units that are the ratio: (the units of y)/(the units of x_i). So, if y has arbitrary BOLD units, then the coefficient/weight will have that arbitrary unit in the numerator, too. This would mean that the overall units of the coefficient would not be easily interpretable or comparable across subjects (and probably not even across the brain).
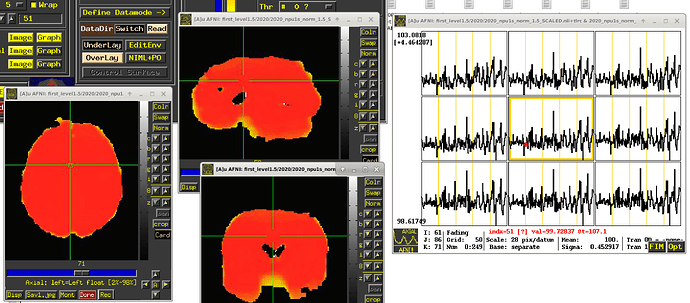
To avoid this scenario, we typically suggest that the BOLD time series should be scaled at each voxel to have meaningful units of "BOLD percent signal change". That is what the inclusion of the "scale" block in afni_proc.py will accomplish. Each voxel is scaled by its own mean to have a mean value of 100, and the fluctuations represent percent changes; so, now, seeing a value of "102" in a subject would be easily interpretable as having a 2% BOLD increase. This is described more in this paper (esp. section [C-3] Presenting results: including the effect estimates.):
and also the case of why this is generally important for the field to appropriately scale data and report/show those values here (note that different software have different default scalings, and so knowing what each one does is important; again, it is hard not to see how local/voxelwise scaling is most appropriate for FMRI):
So, the TL;DR: if you haven't scaled your data as part of processing prior to regression, the coefficients won't have interpretable units. Using afni_proc.py would help do this directly with processing, by including the scale block, which is in many of the examples.
--pt