afni --version
Precompiled binary macos_13_ARM: Apr 1 2026 (Version AFNI_26.0.11 'Pupienus Maximus')
Hi Afni Experts,
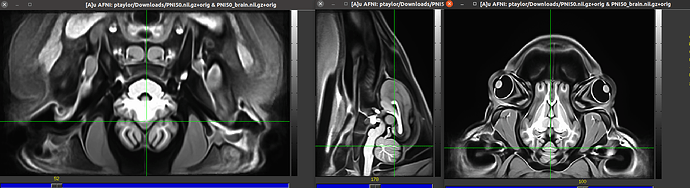
I am currently trying to use @animal_warper to align a dataset to a template, save the warp and inverse warp, apply those transforms to follower datasets like atlases, segmentations, and masks, and do skull stripping by warping a template brain mask back to native space. I looked at both the datasets in ITK snap and both the native and template centers are way apart. So I used itksnap to clisely register them to have the same center as the template space and use animal warper. But everytime i run following code, I get the following error. Could some please shine light on why this is happening. I perfromed this before brining the centers of both the datasets close and wide apart.
#!/usr/bin/env bash
set -euo pipefail
# ============================================================
# Usage:
# bash step1_aw_manual_followers.sh /path/to/sub-001
# ============================================================
SUBJ_DIR="$1"
IN_DIR="${SUBJ_DIR}/input"
OUT_DIR="${SUBJ_DIR}/proc/step1_aw_manual"
mkdir -p "${OUT_DIR}"
# ------------------------------------------------------------
# USE NATIVE SUBJECT IMAGES HERE, NOT T1toPNI50 / T2toPNI50
# ------------------------------------------------------------
T1="${IN_DIR}/t1.nii.gz"
T2="${IN_DIR}/t2.nii.gz"
TEMPLATE_T1="/Volumes/work/Python_projects/Volumetric_analysis/Project_test/T1_template_PIGI50/PNI50.nii.gz"
TEMPLATE_BRAINMASK="/Volumes/work/Python_projects/Volumetric_analysis/Project_test/T1_template_PIGI50/mask_filled.nii.gz"
TEMPLATE_GM="/Volumes/work/Python_projects/Volumetric_analysis/Project_test/T1_template_PIGI50/gm.nii.gz"
TEMPLATE_WM="/Volumes/work/Python_projects/Volumetric_analysis/Project_test/T1_template_PIGI50/wm.nii.gz"
TEMPLATE_CSF="/Volumes/work/Python_projects/Volumetric_analysis/Project_test/T1_template_PIGI50/csf.nii.gz"
[[ -f "${T1}" ]] || { echo "ERROR: Missing ${T1}"; exit 1; }
[[ -f "${TEMPLATE_T1}" ]] || { echo "ERROR: Missing ${TEMPLATE_T1}"; exit 1; }
[[ -f "${TEMPLATE_BRAINMASK}" ]] || { echo "ERROR: Missing ${TEMPLATE_BRAINMASK}"; exit 1; }
[[ -f "${TEMPLATE_GM}" ]] || { echo "ERROR: Missing ${TEMPLATE_GM}"; exit 1; }
[[ -f "${TEMPLATE_WM}" ]] || { echo "ERROR: Missing ${TEMPLATE_WM}"; exit 1; }
[[ -f "${TEMPLATE_CSF}" ]] || { echo "ERROR: Missing ${TEMPLATE_CSF}"; exit 1; }
echo "=== Step 1 manual AFNI workaround ==="
# ------------------------------------------------------------
# 1. Reorient native images
# ------------------------------------------------------------
3dresample -orient RAI -input "${T1}" -prefix "${OUT_DIR}/T1_RAI.nii.gz"
HAS_T2=0
if [[ -f "${T2}" ]]; then
3dresample -orient RAI -input "${T2}" -prefix "${OUT_DIR}/T2_RAI.nii.gz"
HAS_T2=1
fi
cd "${OUT_DIR}"
# ------------------------------------------------------------
# 2. Run @animal_warper WITHOUT follower datasets
# We only want the registration products here
# ------------------------------------------------------------
@animal_warper \
-input "${OUT_DIR}/T1_RAI.nii.gz" \
-input_abbrev subjT1 \
-base "${TEMPLATE_T1}" \
-base_abbrev PNI50 \
-skullstrip "${TEMPLATE_BRAINMASK}" \
-outdir aw_results \
-ok_to_exist
# ------------------------------------------------------------
# 3. Find the actual qwarp output written by auto_warp
# ------------------------------------------------------------
QWARP=$(find aw_results -path "*awpy_*" -name "anat.*.qw_WARP.nii*" | head -n 1 || true)
if [[ -z "${QWARP}" ]]; then
echo "ERROR: Could not find qwarp output."
find aw_results -type f | sort
exit 1
fi
echo "Found qwarp file:"
echo " ${QWARP}"
# ------------------------------------------------------------
# 4. Find affine matrix produced by @animal_warper
# ------------------------------------------------------------
AFF_MAT=$(find aw_results/intermediate -name "*_al2std_mat.aff12.1D" | head -n 1 || true)
if [[ -z "${AFF_MAT}" ]]; then
echo "ERROR: Could not find affine matrix."
find aw_results/intermediate -type f | sort
exit 1
fi
echo "Found affine matrix:"
echo " ${AFF_MAT}"
AFF_INV="aw_results/intermediate/subjT1_inv_al2std_mat.aff12.1D"
cat_matvec -ONELINE "${AFF_MAT}" -I > "${AFF_INV}"
# ------------------------------------------------------------
# 5. Build explicit inverse nonlinear warp from the REAL qwarp
# ------------------------------------------------------------
QWARP_INV="aw_results/intermediate/subjT1_qwarpINV.nii.gz"
3dNwarpCat -prefix "${QWARP_INV}" "INV(${QWARP})"
# ------------------------------------------------------------
# 6. Compose full warps that mirror AFNI's intended logic
# native -> template
# template -> native
# ------------------------------------------------------------
OSH2BASE="aw_results/intermediate/subjT1_native2template_WARP.nii.gz"
BASE2OSH="aw_results/intermediate/subjT1_template2native_WARP.nii.gz"
3dNwarpCat \
-warp1 "${QWARP}" \
-warp2 "${AFF_MAT}" \
-prefix "${OSH2BASE}"
3dNwarpCat \
-warp1 "${AFF_INV}" \
-warp2 "${QWARP_INV}" \
-space NO-DSET \
-prefix "${BASE2OSH}"
# ------------------------------------------------------------
# 7. Apply template masks back into native space
# ------------------------------------------------------------
3dNwarpApply \
-nwarp "${BASE2OSH}" \
-source "${TEMPLATE_BRAINMASK}" \
-master "${OUT_DIR}/T1_RAI.nii.gz" \
-interp NN \
-prefix aw_results/brainmask_in_native.nii.gz
3dNwarpApply \
-nwarp "${BASE2OSH}" \
-source "${TEMPLATE_GM}" \
-master "${OUT_DIR}/T1_RAI.nii.gz" \
-interp NN \
-prefix aw_results/GM_in_native.nii.gz
3dNwarpApply \
-nwarp "${BASE2OSH}" \
-source "${TEMPLATE_WM}" \
-master "${OUT_DIR}/T1_RAI.nii.gz" \
-interp NN \
-prefix aw_results/WM_in_native.nii.gz
3dNwarpApply \
-nwarp "${BASE2OSH}" \
-source "${TEMPLATE_CSF}" \
-master "${OUT_DIR}/T1_RAI.nii.gz" \
-interp NN \
-prefix aw_results/CSF_in_native.nii.gz
# ------------------------------------------------------------
# 8. Threshold / binarize
# ------------------------------------------------------------
3dcalc -a aw_results/brainmask_in_native.nii.gz -expr 'step(a-0.5)' -prefix aw_results/brainmask_in_native_bin.nii.gz
3dcalc -a aw_results/GM_in_native.nii.gz -expr 'step(a-0.5)' -prefix aw_results/GM_in_native_bin.nii.gz
3dcalc -a aw_results/WM_in_native.nii.gz -expr 'step(a-0.5)' -prefix aw_results/WM_in_native_bin.nii.gz
3dcalc -a aw_results/CSF_in_native.nii.gz -expr 'step(a-0.5)' -prefix aw_results/CSF_in_native_bin.nii.gz
# ------------------------------------------------------------
# 9. Skull-stripped native T1
# ------------------------------------------------------------
3dcalc \
-a "${OUT_DIR}/T1_RAI.nii.gz" \
-b aw_results/brainmask_in_native_bin.nii.gz \
-expr 'a*step(b)' \
-prefix aw_results/T1_native_brain.nii.gz
# ------------------------------------------------------------
# 10. Final tissue label map
# 1 = CSF
# 2 = GM
# 3 = WM
# ------------------------------------------------------------
3dcalc \
-a aw_results/CSF_in_native_bin.nii.gz \
-b aw_results/GM_in_native_bin.nii.gz \
-c aw_results/WM_in_native_bin.nii.gz \
-expr '3*step(c)+2*step((1-step(c))*b)+1*step((1-step(c))*(1-step(b))*a)' \
-prefix aw_results/tissue_labels_native.nii.gz
echo
echo "=== Done ==="
echo "Check:"
echo " aw_results/T1_native_brain.nii.gz"
echo " aw_results/brainmask_in_native_bin.nii.gz"
echo " aw_results/GM_in_native_bin.nii.gz"
echo " aw_results/WM_in_native_bin.nii.gz"
echo " aw_results/CSF_in_native_bin.nii.gz"
echo " aw_results/tissue_labels_native.nii.gz"
echo " ${OSH2BASE}"
echo " ${BASE2OSH}"
echo " ${QWARP}"
The error is the 3dNwrapCat cannot find the follwing file:
** ERROR: Can't open dataset from file 'anat.un.qw_WARP.nii'
** ERROR: Failed to read 3D warp from 'INV(anat.un.qw_WARP.nii)'
** ERROR: Can't compute nonlinear warp from string 'INV(anat.un.qw_WARP.nii) '
** ERROR: EDIT_dset_items[1]: invalid input dataset
** ERROR: Cannot write dataset: it is invalid
++ DONE! Image output:
QC/init_qc_02.input_aff+base.subjT1
#++ auto_warp.py version: 0.06
-- clearing AFNI_COMPRESSOR ...
# Output directory /Volumes/work/Python_projects/Volumetric_analysis/Project_test/sub-001/proc/step1_aw_manual/aw_results/awpy_subjT1_pshft/
#Script is running (command trimmed):
mkdir ./awpy_subjT1_pshft/
cd /Volumes/work/Python_projects/Volumetric_analysis/Project_test/sub-001/proc/step1_aw_manual/aw_results/awpy_subjT1_pshft/
#Script is running (command trimmed):
3dcopy /Volumes/work/Python_projects/Volumetric_analysis/Project_test/sub-001/proc/step1_aw_manual/aw_results/intermediate/subjT1_shft_aff.nii.gz ./anat.nii
++ 3dcopy: AFNI version=AFNI_26.0.11 (Apr 1 2026) [64-bit]
#Script is running (command trimmed):
3dUnifize -GM -input ./anat.nii -prefix ./anat.un.nii
++ 3dUnifize: AFNI version=AFNI_26.0.11 (Apr 1 2026) [64-bit]
+ Pre-processing: ADV...............UW[s120448]Gm
++ Output dataset ./anat.un.nii
++ ===== CPU time = 46.5 sec Elapsed = 9.0
#Script is running (command trimmed):
3dcopy /Volumes/work/Python_projects/Volumetric_analysis/Project_test/T1_template_PIGI50/PNI50.nii.gz ./base.nii
++ 3dcopy: AFNI version=AFNI_26.0.11 (Apr 1 2026) [64-bit]
#Script is running (command trimmed):
3dAttribute DELTA ./anat.un.nii
#Script is running (command trimmed):
3dAttribute DELTA ./base.nii
0.596591 0.596591
#Script is running (command trimmed):
3dinfo -same_grid ./anat.un.nii ./base.nii
#Script is running (command trimmed):
3dresample -inset ./anat.un.nii -prefix ./anat.rwb.nii -rmode Li -master ./base.nii
#++ Aligning /Volumes/work/Python_projects/Volumetric_analysis/Project_test/sub-001/proc/step1_aw_manual/aw_results/awpy_subjT1_pshft/base.nii data to /Volumes/work/Python_projects/Volumetric_analysis/Project_test/sub-001/proc/step1_aw_manual/aw_results/awpy_subjT1_pshft/anat.rwb.nii data
#Script is running (command trimmed):
3dQwarp -prefix ./anat.rwb.qw.nii -blur -3 -3 -workhard:0:2 -maxlev 09 -base ./base.nii -source ./anat.rwb.nii
OMP: Info #276: omp_set_nested routine deprecated, please use omp_set_max_active_levels instead.
++ OpenMP thread count = 15
++ 3dQwarp: AFNI version=AFNI_26.0.11 (Apr 1 2026) [64-bit]
++ Authored by: Zhark the (Hermite) Cubically Warped
++ negative values in base ==> using strict Pearson correlation
++ Dataset final zero-pad: xbot=44 xtop=44 ybot=36 ytop=36 zbot=20 ztop=20 voxels
++ Weightizing the base image: FWHM = 4.5 (vox)
++ +++++++++++ Begin warp optimization: base=/Volumes/work/Python_projects/Volumetric_analysis/Project_test/sub-001/proc/step1_aw_manual/aw_results/awpy_subjT1_pshft/base.nii source=/Volumes/work/Python_projects/Volumetric_analysis/Project_test/sub-001/proc/step1_aw_manual/aw_results/awpy_subjT1_pshft/anat.rwb.nii
++ AFNI warpomatic: 440 x 357 x 190 volume ; autobbox = 41..397 30..323 14..174 [clock= 8s 894ms]
lev=0 1..438 1..355 1..188: [first cost=-0.21020] ..... done [cost:-0.21020==>-0.24715]
lev=1 patch=329x267x141 [clock= 1m 25s 153ms]........:[cost=-0.26590]:........done [cost:-0.24715==>-0.29289 ; 16 patches optimized, 0 skipped, bbox=3:436,3:355,-23:188]
lev=2 patch=247x199x105 [clock= 2m 20s 235ms]...........:[cost=-0.33037]:..........done [cost:-0.29289==>-0.37431 ; 21 patches optimized, 3 skipped, bbox=3:436,3:355,-34:188]
lev=3 patch=185x149x79 [clock= 3m 2s 754ms]............done [cost:-0.37431==>12.47312 ; 32 patches optimized, 4 skipped, bbox=3:436,3:355,-37:188]
lev=4 patch=139x113x59 [clock= 3m 39s 804ms]..............done [cost:12.47312==>17.00923 ; 100 patches optimized, 25 skipped, bbox=3:436,3:355,-35:188]
lev=5 patch=103x85x45 [clock= 4m 27s 791ms]................done [cost:17.00923==>9.88767 ; 202 patches optimized, 50 skipped, bbox=3:436,3:355,-30:188]
lev=6 patch=77x63x33 [clock= 5m 19s 796ms]..........................done [cost:9.88767==>9.13047 ; 599 patches optimized, 211 skipped, bbox=3:436,3:355,-26:188]
lev=7 patch=59x47x25 [clock= 6m 29s 410ms].........................................done [cost:9.13047==>5.88609 ; 1335 patches optimized, 537 skipped, bbox=3:436,3:355,-20:188]
lev=8 patch=43x35x19 [clock= 8m 2s 323ms].........................................................done [cost:5.88609==>5.05206 ; 3317 patches optimized, 1596 skipped, bbox=3:436,3:355,-18:188]
lev=9 patch=33x27x15 [clock= 10m 30s 318ms]..............................................................................................done [cost:5.05206==>4.93081 ; 7178 patches optimized, 3470 skipped, bbox=3:436,3:355,-16:188]
++ ====== total number of parameters 'optimized' = 153750
+ initial unpenalized cost = -0.210196
+ final unpenalized cost = -0.41405
+ final penalized cost = 4.93081
++ Output dataset ./anat.rwb.qw.nii
++ Output dataset ./anat.rwb.qw_WARP.nii
++ ===== CPU time = 4266.7 sec clock time = 14m 48s 783ms
#++ Applying warps to /Volumes/work/Python_projects/Volumetric_analysis/Project_test/sub-001/proc/step1_aw_manual/aw_results/intermediate/subjT1_shft_aff.nii.gz
#Script is running (command trimmed):
3dNwarpApply -nwarp ./anat.rwb.qw_WARP.nii -master ./base.nii -source /Volumes/work/Python_projects/Volumetric_analysis/Project_test/sub-001/proc/step1_aw_manual/aw_results/intermediate/subjT1_shft_aff.nii.gz -prefix ./subjT1_shft_aff.aw.nii
++ 3dNwarpApply: AFNI version=AFNI_26.0.11 (Apr 1 2026) [64-bit]
++ Authored by: Zhark the Warped
++ -master dataset is './base.nii'
++ opened source dataset '/Volumes/work/Python_projects/Volumetric_analysis/Project_test/sub-001/proc/step1_aw_manual/aw_results/intermediate/subjT1_shft_aff.nii.gz'
++ Processing -nwarp
++ Warping:.Z
++ Output dataset ./subjT1_shft_aff.aw.nii
++ total CPU time = 33.9 sec Elapsed = 2.7
#++ Saving history
#Script is running (command trimmed):
3dNotes -h "auto_warp.py -overwrite -base \
/Volumes/work/Python_projects/Volumetric_analysis/Project_test/T1_template_PIGI50/PNI50.nii.gz \
-affine_input_xmat ID -qworkhard 0 2 -input \
intermediate/subjT1_shft_aff.nii.gz -output_dir awpy_subjT1_pshft \
-qw_opts -maxlev 09" \
./subjT1_shft_aff.aw.nii
++ 3dNwarpCat: AFNI version=AFNI_26.0.11 (Apr 1 2026) [64-bit]
++ Authored by: Zhark the Warper
** ERROR: Can't open dataset from file 'anat.un.qw_WARP.nii'
** ERROR: Failed to read 3D warp from 'INV(anat.un.qw_WARP.nii)'
** ERROR: Can't compute nonlinear warp from string 'INV(anat.un.qw_WARP.nii) '
** ERROR: EDIT_dset_items[1]: invalid input dataset
** ERROR: Cannot write dataset: it is invalid
Fatal Signal 11 (SIGSEGV) received
3dNwarpCat
Bottom of Debug Stack
** AFNI version = AFNI_26.0.11 Compile date = Apr 1 2026
** [[Precompiled binary macos_13_ARM: Apr 1 2026]]
** Program Death **
** If you report this crash to the AFNI message board,
** please copy the error messages EXACTLY, and give
** the command line you used to run the program, and
** any other information needed to repeat the problem.
** You may later be asked to upload data to help debug.
** Crash log is appended to file /Users/myru/.afni.crashlog
++ 3drefit: AFNI version=AFNI_26.0.11 (Apr 1 2026) [64-bit]
++ Authored by: RW Cox
** ERROR: Can't open dataset anat.un.qw_WARPINV.nii
++ 3drefit processed 0 datasets
++ 3dcopy: AFNI version=AFNI_26.0.11 (Apr 1 2026) [64-bit]
3dcopy: No match.
** ERROR: program failed (autowarp cp)```
The program ended and I am not sure what the case of failure is.

Thank you for your help in advance.
Pavan.