Precompiled binary macos_13_ARM: Jul 7 2025 (Version AFNI_25.2.03 'Gordian I')
Hi there,
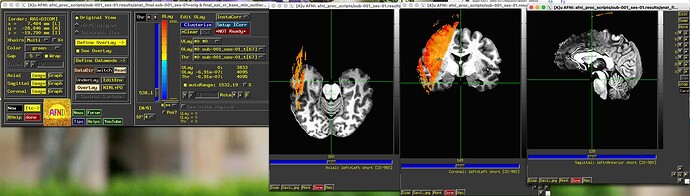
I just ran a pre-processing pipeline on some task fMRI data. The alignment step failed, in that the epi and anat are not well aligned. I was just testing it on one participant.
Underlay below: anat_final.sub-001_ses-01
Overlay: final_epi_vr_base_min_outlier
#!/bin/bash
# Set up participant details
sub=sub-001
ses=ses-01
# Set up directories
base_dir=/Users/uqhdemp1/Library/CloudStorage/OneDrive”
RDM_base_dir=${base_dir}
script_dir=${base_dir}/Scripts/afni_proc_scripts
extension=".nii.gz"
anat_raw_dir=${RDM_base_dir}/Data_from_RDM/bids/${sub}/${ses}/anat
anat_raw=${sub}_${ses}_acq-UNIDEN_run-1_T1w
data_dir=${RDM_base_dir}/Data_from_RDM/bids/${sub}/${ses}/func
data_base=${sub}_${ses}_task-PHASE
surf_dir=${RDM_base_dir}/Data_from_RDM/bids/derivatives/fastsurfer/output/${sub}/${ses}/SUMA
blur_size="2"
afni_proc.py \
-subj_id ${sub}_${ses} \
-dsets "$data_dir/${data_base}"*"${extension}" \
-blocks tcat despike align volreg surf blur scale \
-copy_anat $anat_raw_dir/${anat_raw}${extension} \
-anat_has_skull yes \
-surf_anat $surf_dir/${sub}_${ses}_SurfVol.nii \
-surf_spec $surf_dir/${sub}_${ses}_?h.spec \
-tcat_remove_first_trs 0 \
-align_opts_aea -giant_move \
-partial_coverage \
-cost lpc \
-align_unifize_epi local \
-volreg_align_e2a \
-volreg_align_to MIN_OUTLIER \
-volreg_post_vr_allin yes \
-volreg_pvra_base_index MIN_OUTLIER \
-volreg_interp -Fourier \
-volreg_warp_final_interp wsinc5 \
-volreg_compute_tsnr yes \
-blur_size $blur_size \
-html_review_style pythonic
I tried to see if a different cost function might improve things. Code below:
(note: I just realised I was using epi2anat, which is not what I want, but the principles should still hold right?)
#!/bin/bash
subj=sub-001_ses-01
basedir="/Users/uqhdemp1/Library/CloudStorage/OneDrive"
data_loc_proc=$basedir/MND/Scripts/afni_proc_scripts/${subj}.results
data_loc=${data_loc_proc}-align_tests
anat=${subj}_acq-UNIDEN_run-1_T1w+orig
epi_unif=vr_base_min_outlier_unif+orig
# Voxel size to apply warps to
set mast_dxyz = 0.75
cd $data_loc
align_epi_anat.py -epi2anat \
-anat $anat \
-anat_has_skull yes \
-suffix _al_junk \
-epi $epi_unif \
-epi_base 0 \
-epi_strip 3dAutomask \
-giant_move \
-partial_coverage \
-cost lpc+ZZ -multi_cost lpc lpa \
-volreg off -tshift off
The lpc produced (what looks like) the same results as in my afni_proc.py script. The other two (lpa, lpc+ZZ) were worse.
I tried lpc on its own, without giant move (didn’t work), then with big move (didn’t work), and with ginormous move (didn’t work).
I tried no epi skull stripping, and that did not work either.
-epi_strip None \
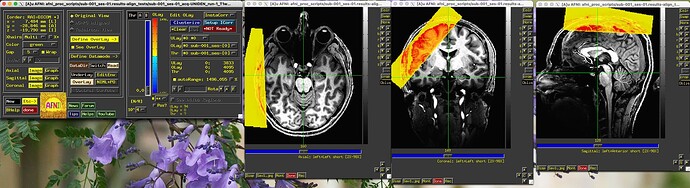
Interestingly, I tried to align the raw epi to the anat, using the same run/ sub-brick that the min_outlier was taken from, run 01, 67 sub-brick. The lpc and lpa worked… I This is good, but not really helpful in fixing my afni_proc.py pipeline.
Underlay below: sub-001_ses-01-acq-UNIDEN_run-1_T1W
Overlay: sub-001_ses-01_task-PHASE1_bold_67_al_junc_lpc
#!/bin/bash
subj=sub-001_ses-01
basedir="/Users/uqhdemp1/Library/CloudStorage/OneDrive"
data_loc_proc=$basedir/MND/Scripts/afni_proc_scripts/${subj}.results
data_loc=${data_loc_proc}-align_tests
3dbucket ${subj}_task-PHASE1_bold.nii.gz[0] ${subj}_task-PHASE1_bold_0+orig
anat=${subj}_acq-UNIDEN_run-1_T1w+orig
epi=${subj}_task-PHASE1_bold_0+orig
apply_only=0
# Voxel size to apply warps to
set mast_dxyz = 0.75
cd $data_loc
align_epi_anat.py -epi2anat \
-anat $anat \
-anat_has_skull yes \
-suffix _al_junk \
-epi $epi \
-epi_base 0 \
-epi_strip 3dAutomask \
-partial_coverage \
-ginormous_move \
-cost lpc \
-volreg off -tshift off
I have also previously run the same preprocessing with my input anatomical being a skull stripped anatomical from HDBet. This worked fine too. I was going to try this again, but need to download HDbet and remake the skull stripped images.
As an extra side question – I read in lots of the documentation that lpc+ZZ is the most robust, but it never seems to work with my data. Any idea why? I always use high resolution (0.5-075 anatomicals, and 0.8ish epi partial coverage)
Thank you so much for your help!
H